AncestryDNA region data
Ancestry's chromsome painter shows 'the regions' they associate the test taker's DNA.
Their current regions:
Ancestry also updates it's regions over time. The last update was 10 Oct 2025.
How can I use this information to support my research?
The information might:
- show locations of interest for segments where there is no MRCA mapped.
- be evidence that supports or contradict conclusions where MRCA is uncertain.
Can I add the region data to GDAT's segment map?
2026-02-08
Bottom line up front
I didn't get it to work this time, so reverted to DNA Painter. Maybe next time.
I am using DNAPainter, but wanted test how I might add the information to GDAT for easier referencing.
- Created the csv file following DNA Painter's instructions.
- DNA Painter csv filename: ~#~2026_01_30_Ancestry_acp-segments.csv~#~
- There is no cM totals. Later realised GDAT needs these totals.
- Tested adding the region names to GDAT as relatives. (This worked).
- Created a csv file of the list of unique region names:
- Added 'segments_as_relatives' to the filename.
- Copied the group column from the DNA Painter csv file and removed duplicates.
- Created groupID column with unique identifier for each region name (as a substitute for the relative primary key in GDAT).
- Added the region names as relatives to GDAT
- In GDAT, create an Import Template.
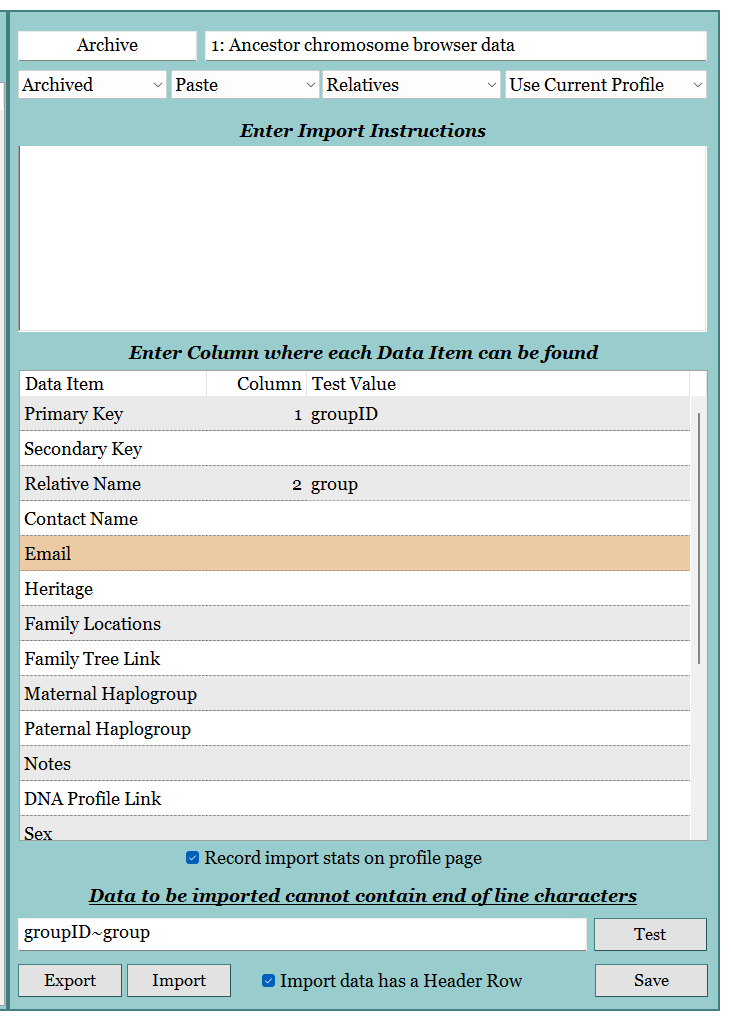
- Forget to add instructions, so had to start over again.
- In GDAT, create an Import Template.
- Created a csv file of the list of unique region names:
- Tested adding chromosome data for GDAT to map (I didn't get this to work)
- In the DNAPainter csv file, added a groupID column and used an XLOOKUP function to add the groupID from the segments_as_relatives csv file.
- In GDAT, created an Import Template.
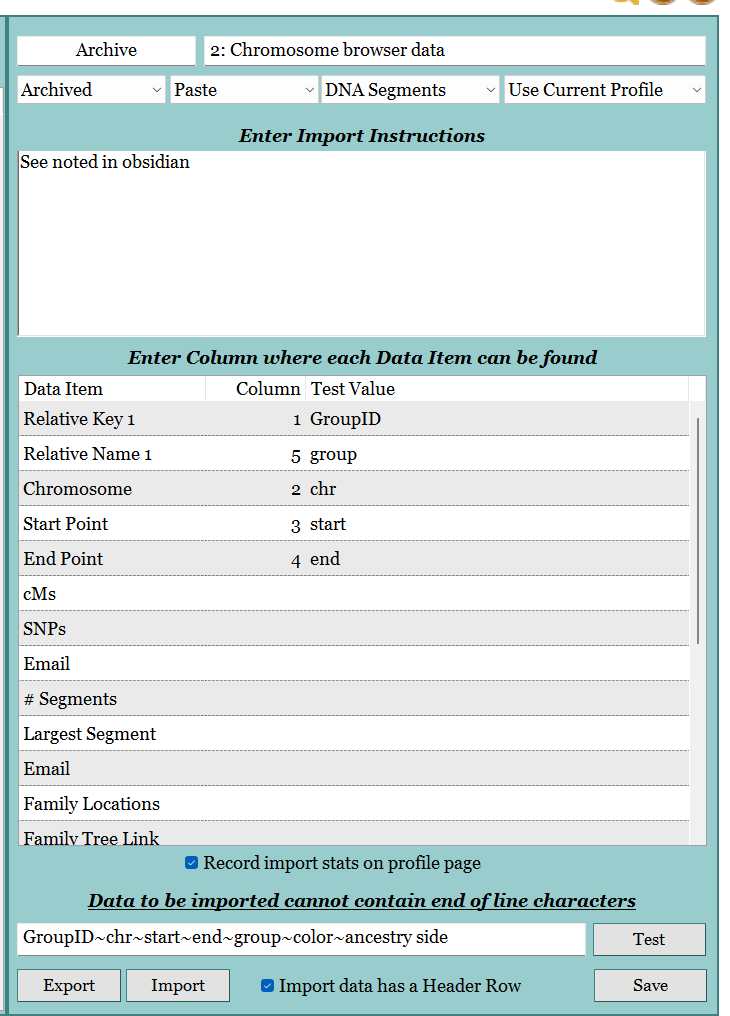
- Original import attempt failed as the cMs was below the minimum required.
- To fix, in Preferences (F10), changed minimum cM level to 0

- Second import attempt worked, but 16 segments skipped because 'No Segment Endpoint'
- Reset minimum cM level in Preferences (F10).
- Reviewed relatives page for each group an assigned Paternal or Maternal.
- After looking at the Chromosome Browser (F7), finally realised the impact of cMs=0.
- Tried a fix:
- In Preferences (F10), run Relative Data Routine: 'Recalculate the Total cMs and Segments by Profile'
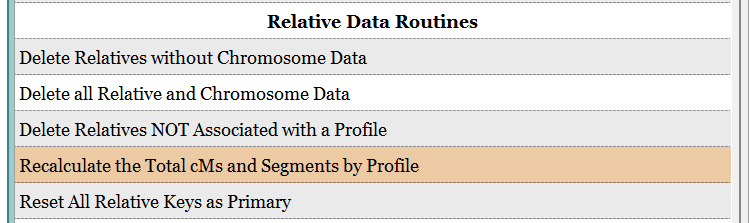
- In Preferences (F10), run Relative Data Routine: 'Recalculate the Total cMs and Segments by Profile'
- Realised that this doesn't make data appear when there is none 🙃 and decided to try again another day.